RNA splicing
In molecular biology, splicing is the editing of the nascent precursor messenger RNA (pre-mRNA) transcript into a mature messenger RNA (mRNA). After splicing, introns are removed and exons are joined together (ligated).
For nuclear-encoded genes, splicing takes place within the nucleus either during or immediately after transcription. For those eukaryotic genes that contain introns, splicing is usually required in order to create a mRNA molecule that can be translated into protein.
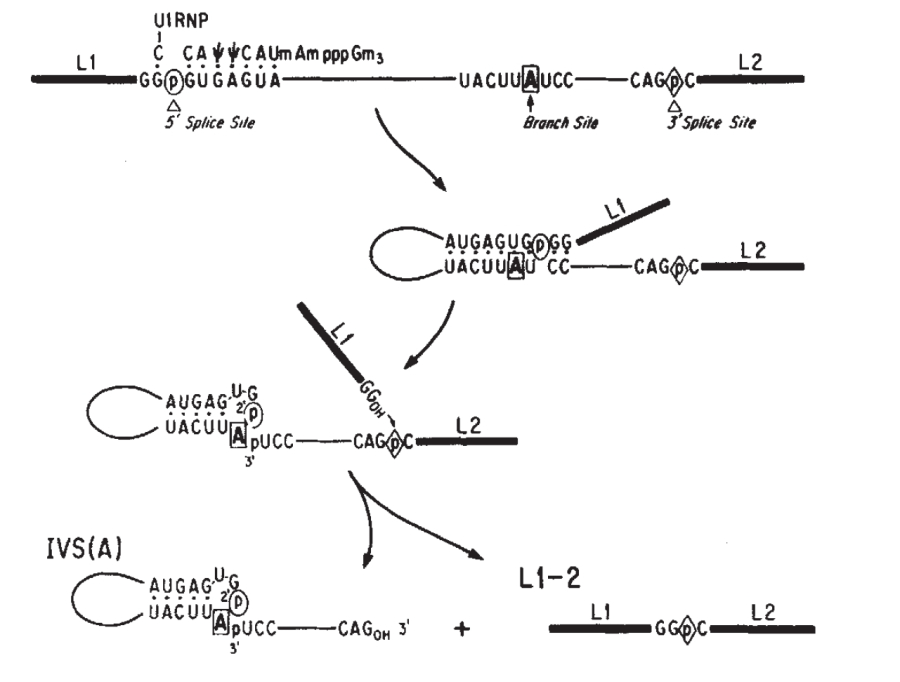
For many eukaryotic introns, splicing is carried out in a series of reactions which are catalyzed by the spliceosome, a complex of snRNPs. Self-splicing introns, or ribozymes capable of catalyzing their own excision from their parent RNA molecule, also exist.
- Group I and II introns perform splicing similar to the spliceosome without requiring any protein. This similarity suggests that Group I and II introns may be evolutionarily related to the spliceosome. Self-splicing may also be very ancient and may have existed in an RNA world present before protein.
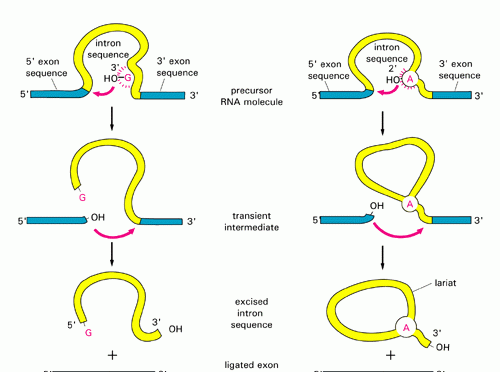
- Two transesterifications characterize the mechanism through which group of introns is spliced:
- 3’OH of a free guanine nucleoside (or one located in the intron) or a nucleotide cofactor (GMP, GDP, GTP) attacks phosphate at the 5′ splice site.
- 3’OH of the 5′ exon becomes a nucleophile and the second transesterification results in the joining of the two exons.

- The mechanism in which group II introns are spliced (two transesterification reaction like group I introns) is as follows:
- The 2’OH of a specific adenosine in the intron attacks the 5′ splice site, thereby forming the lariat
- The 3’OH of the 5′ exon triggers the second transesterification at the 3′ splice site, thereby joining the exons together.